RNAP entries
1.Browse by organism.On the “Organism" page,users can browse the database by clicking the species image of interest.
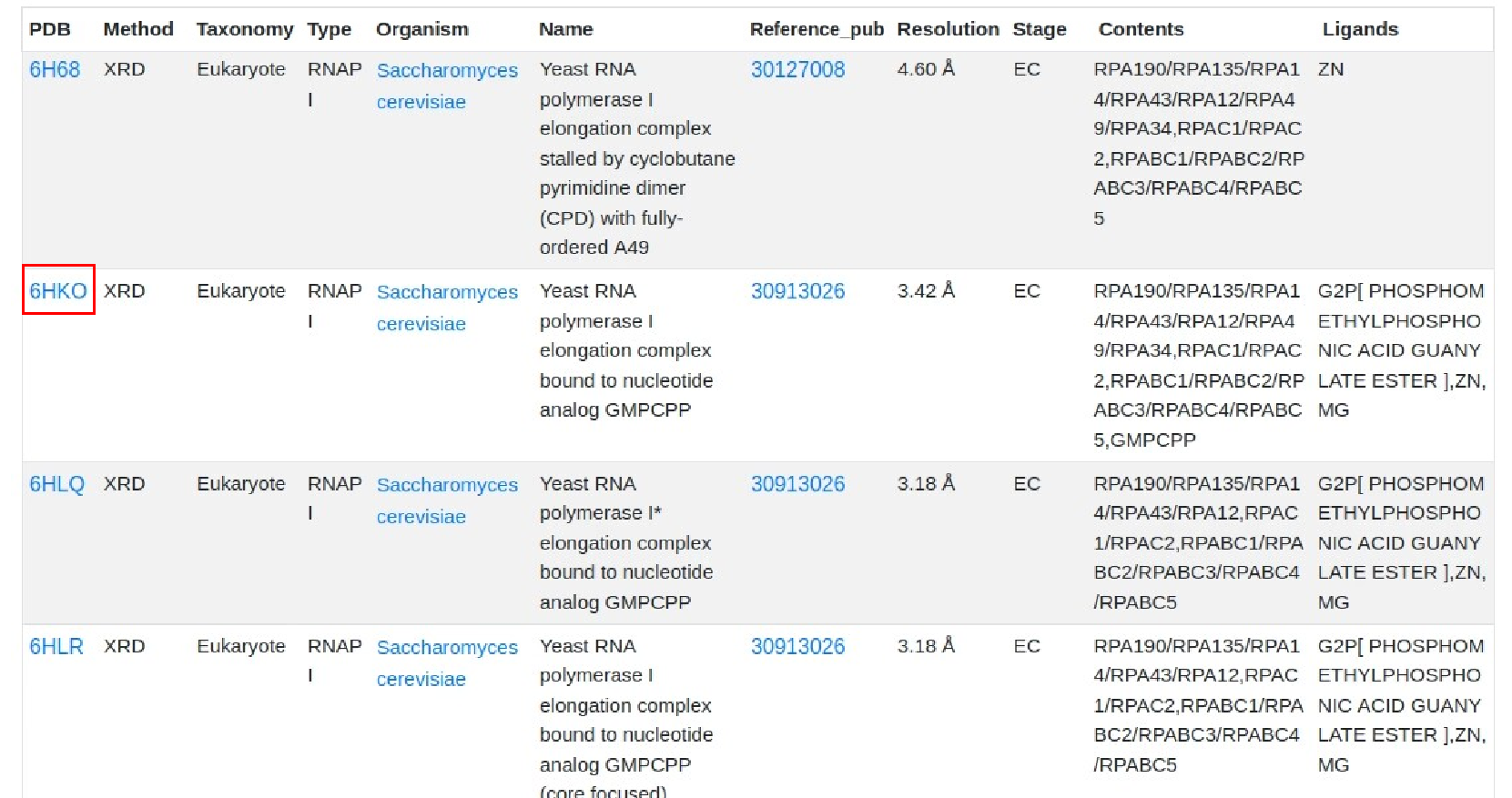
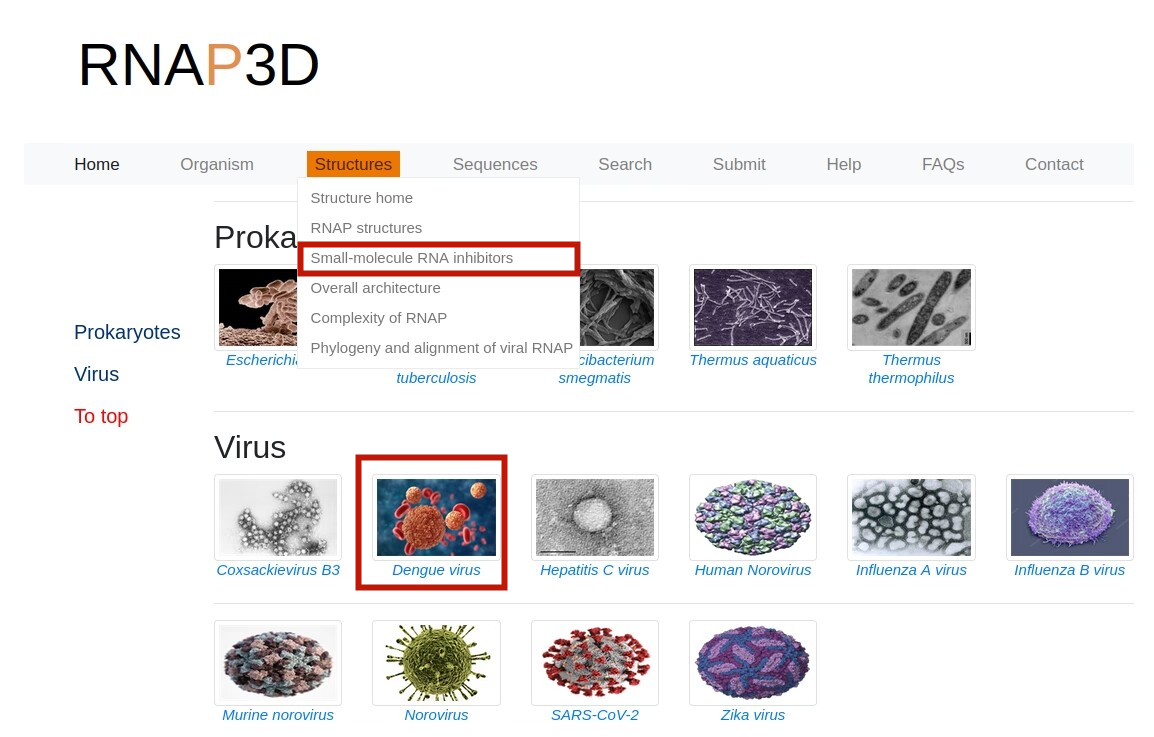
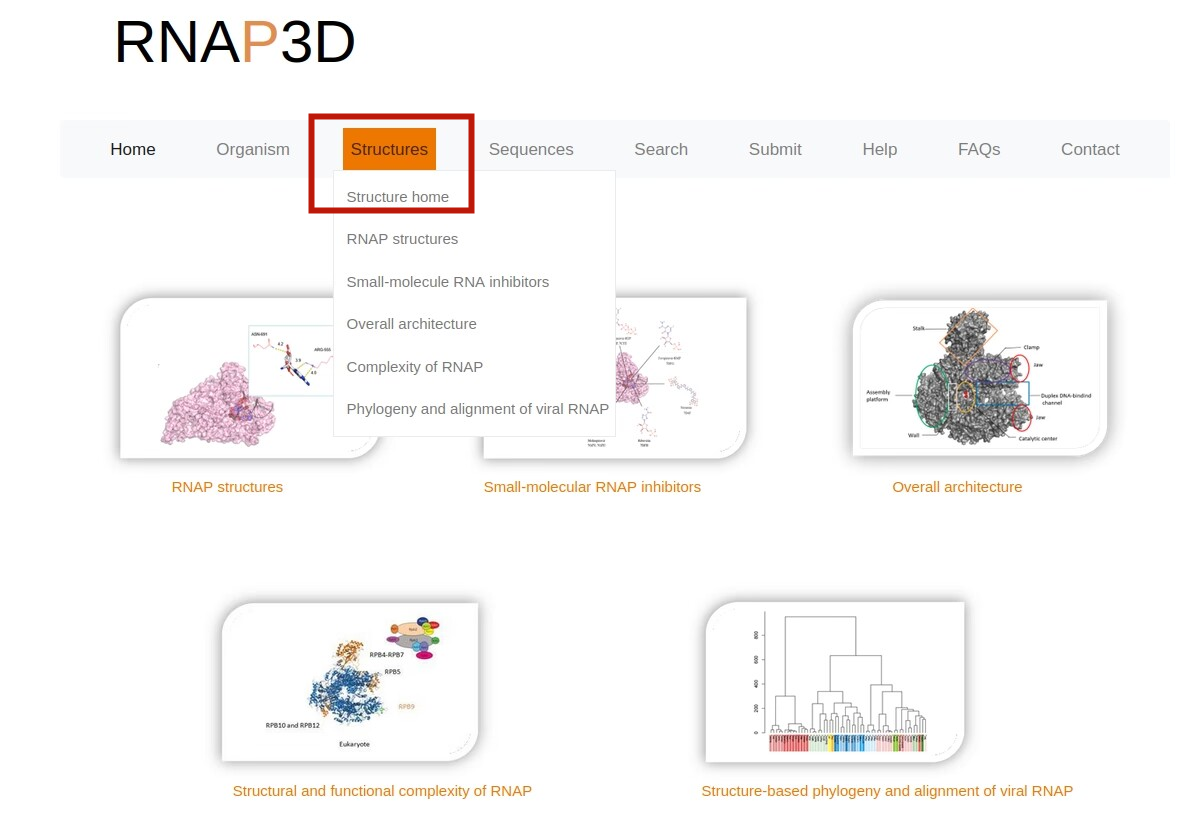
2.Browse by type.On the “Home" page,users can browse the database by clicking the types image of interest.For example, users can retrieve the detailed RNAP I information through the following steps:Eukarytoes -> RNAP I -> PDB.
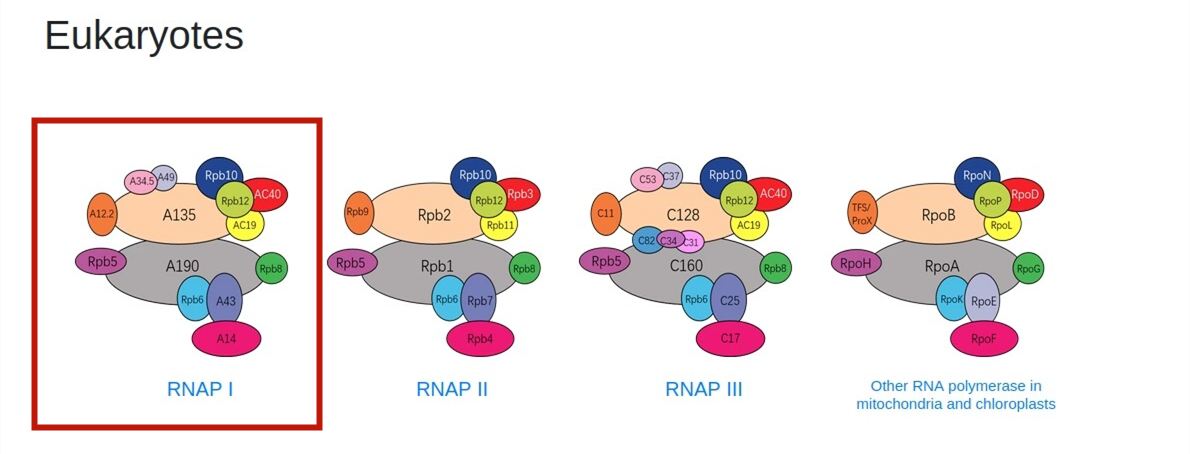
3.Browse by Taxonomy.On the “RNAP structures" page,users can browse the database by clicking the taxonomy image of interest.
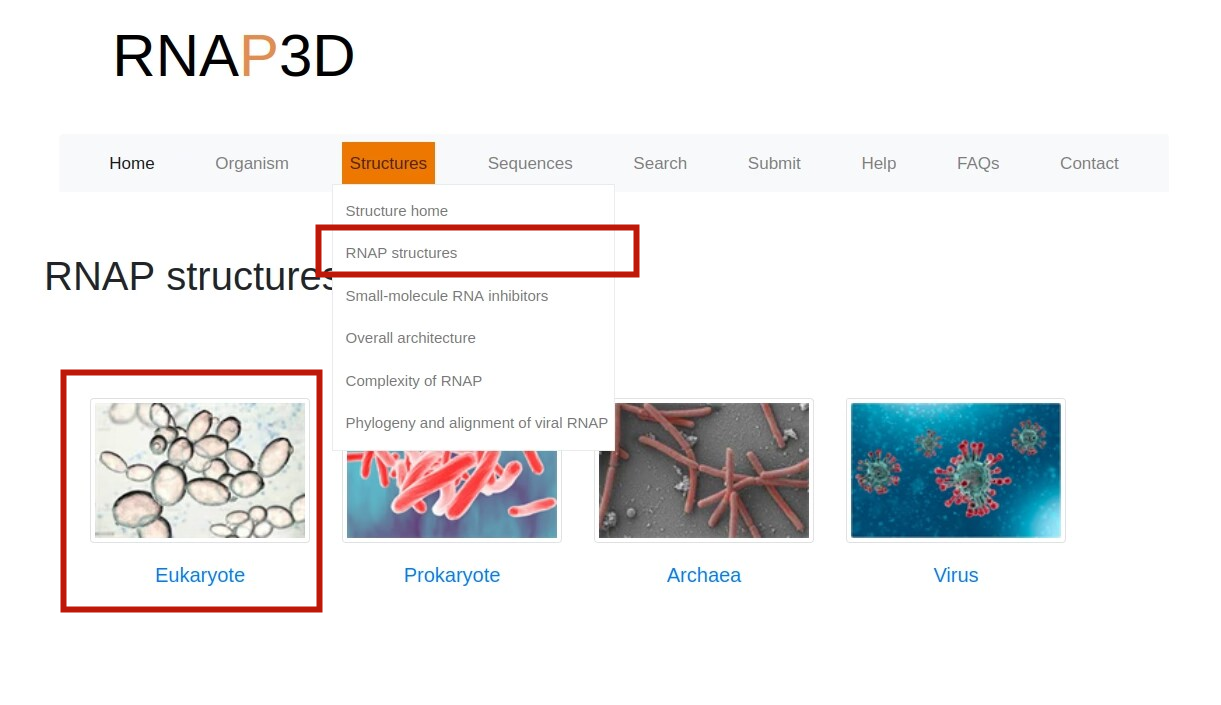
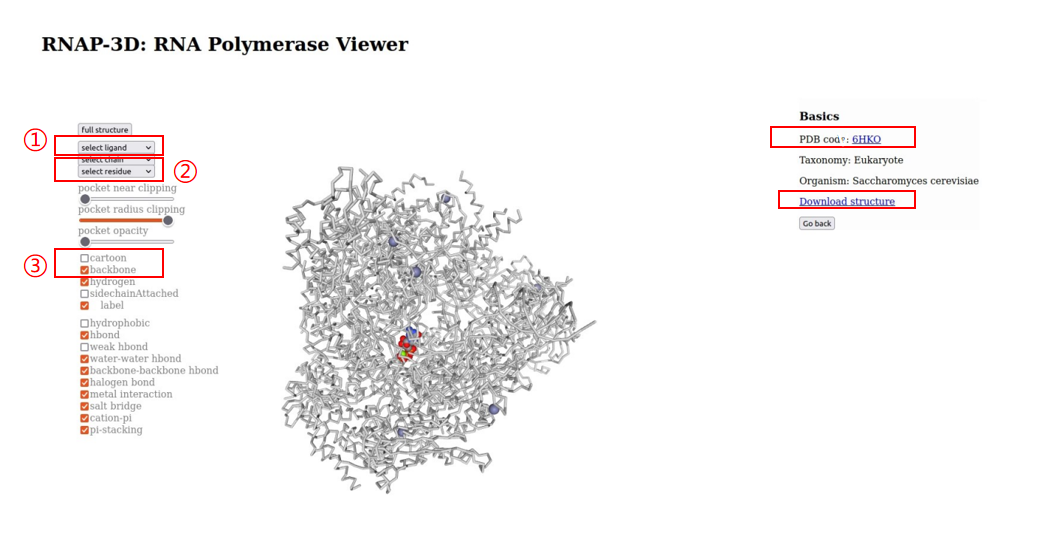
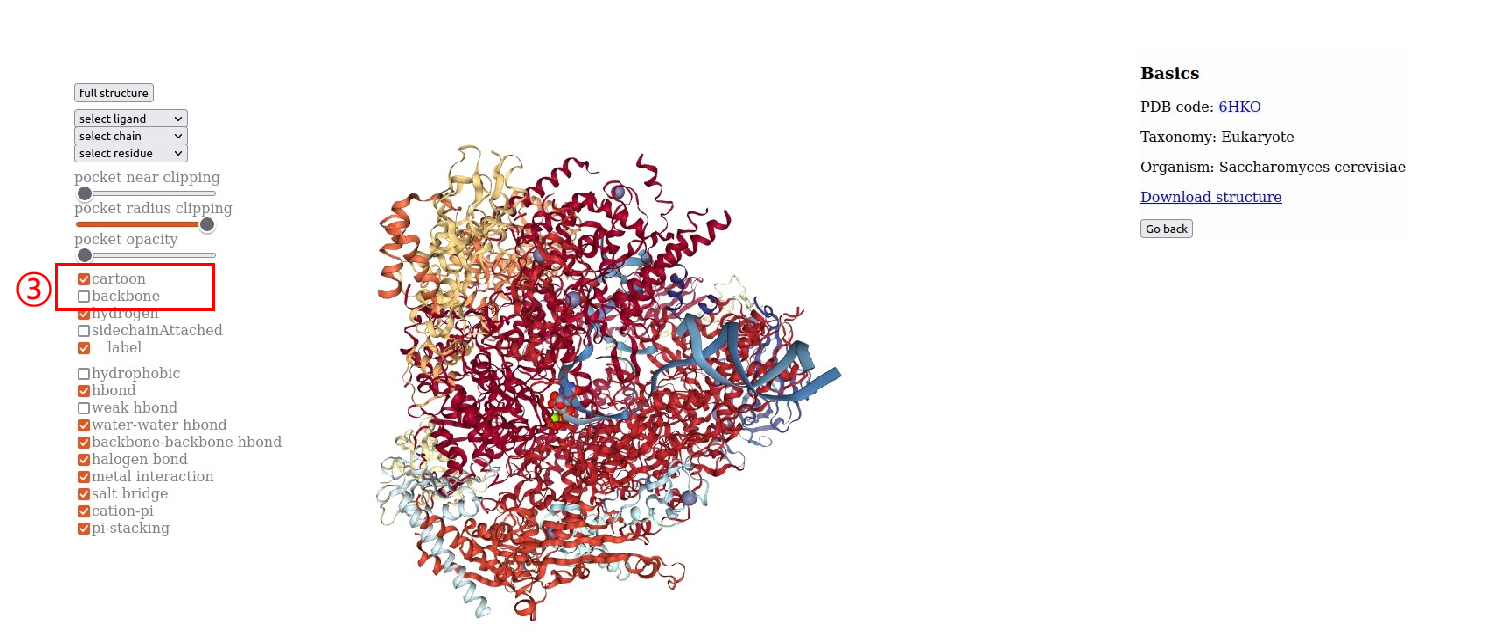
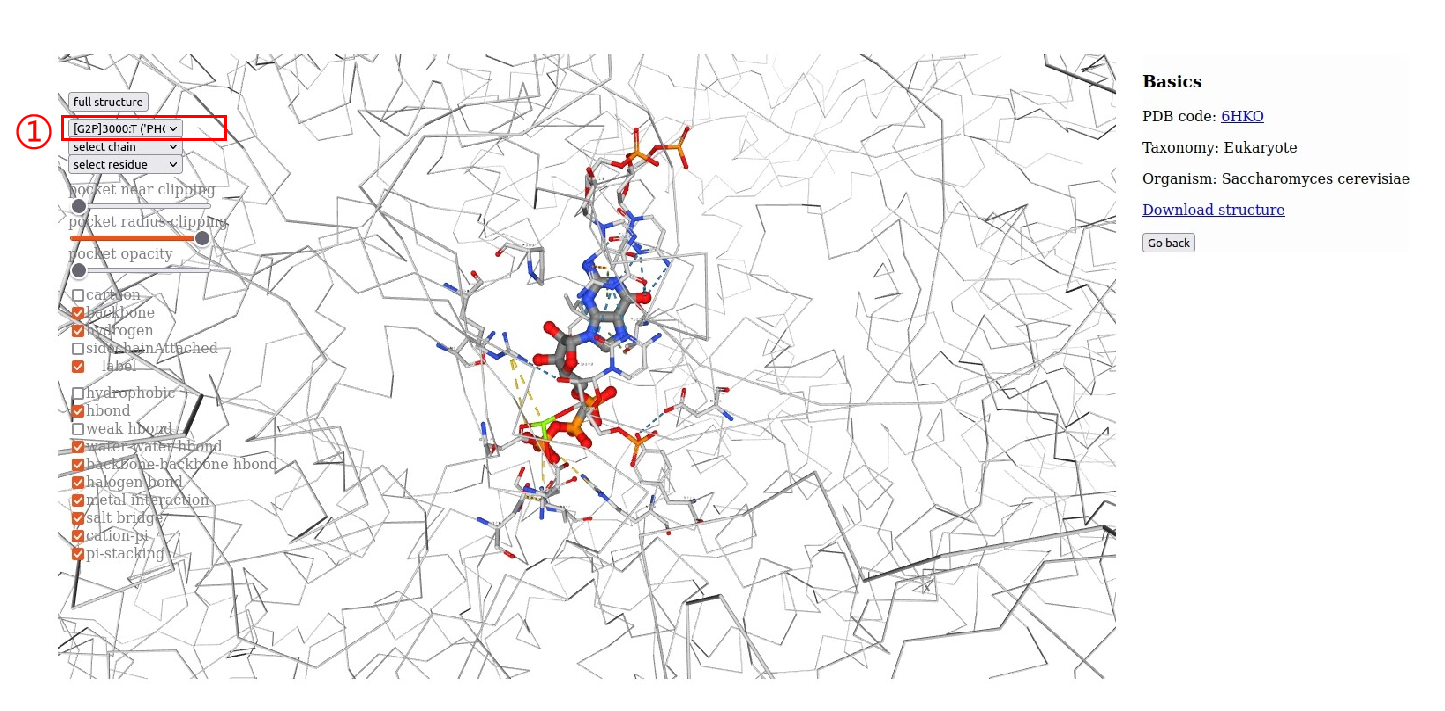
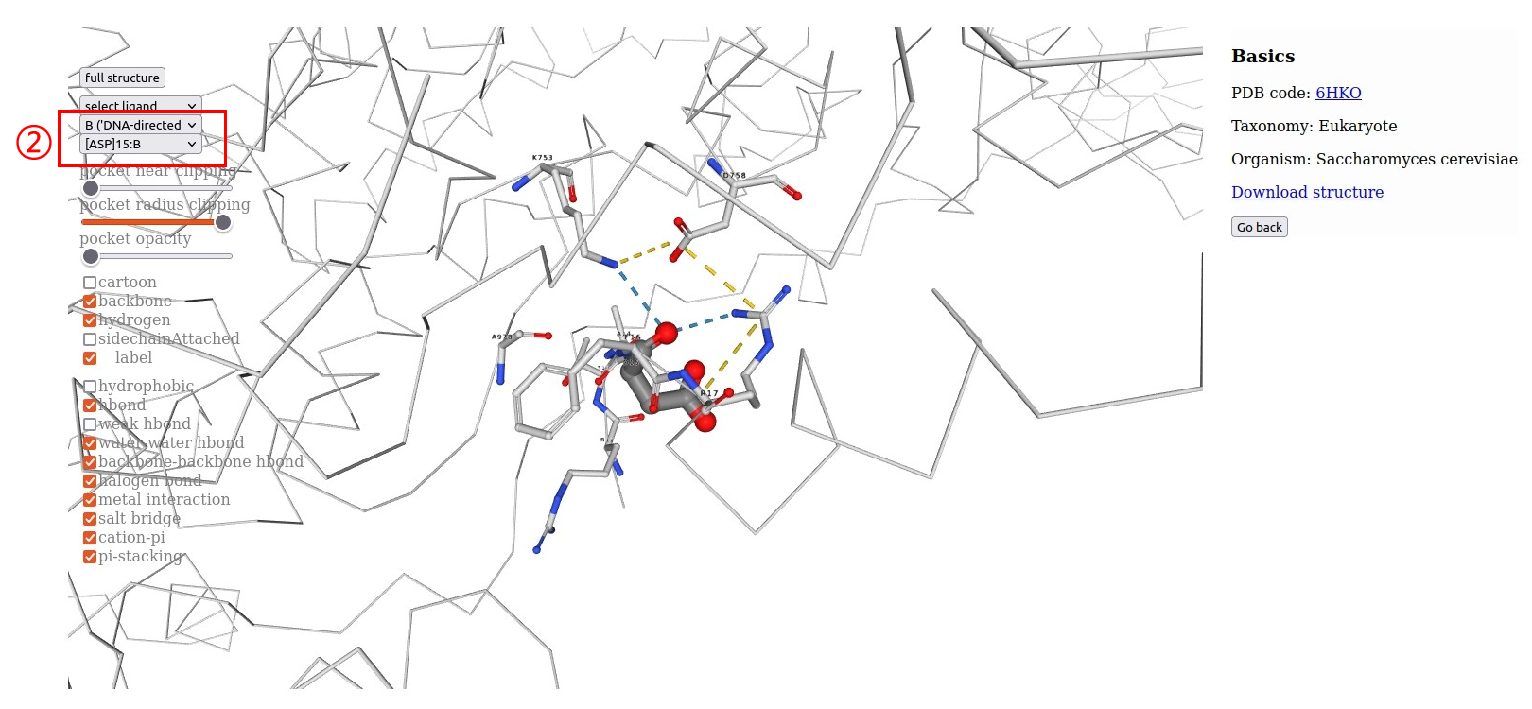
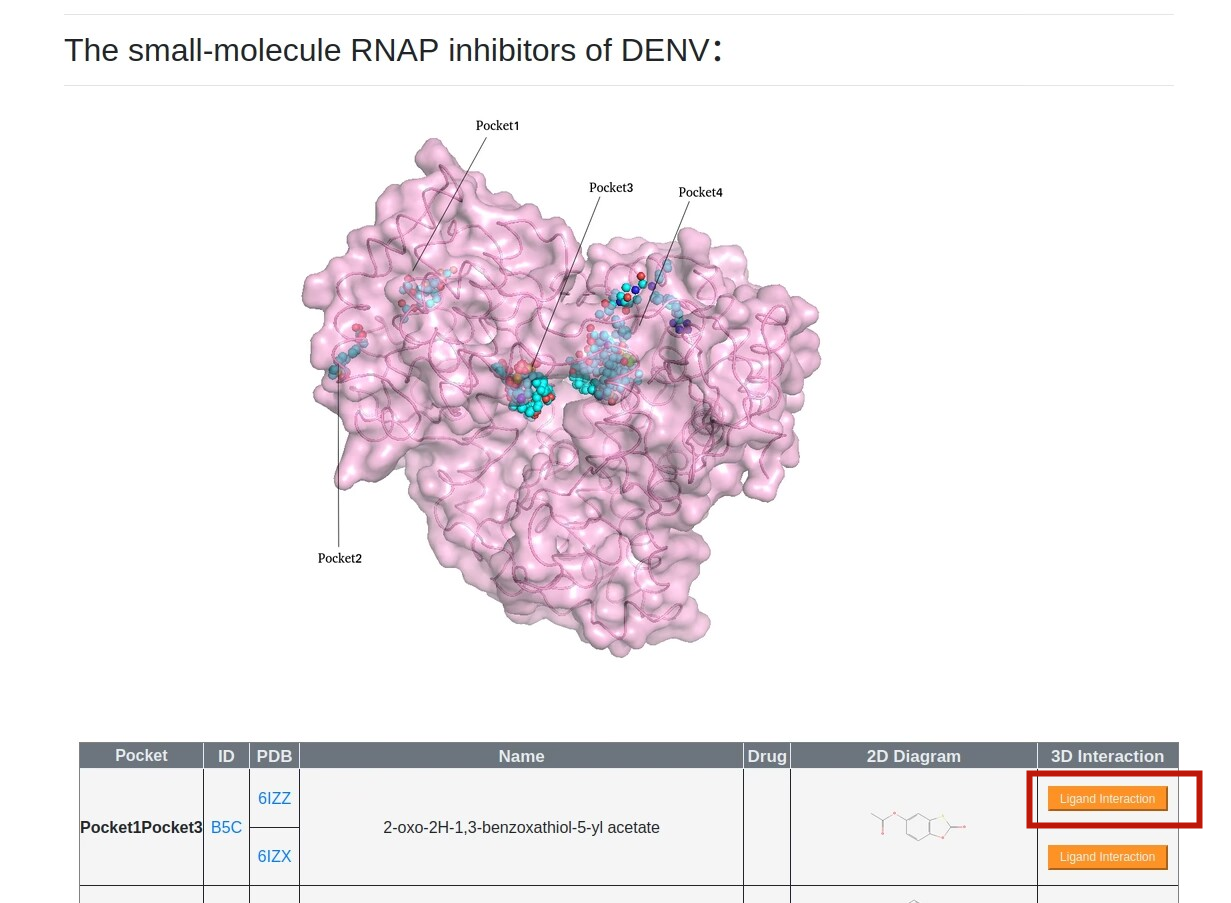
4.RNAP structural information.Users can browse “RNA Polymearase Viewer” page by clicking PDB id,and then display different formats according to their interests. In addition,users can select ligands or residues of different subunits to view their interactions. Users can also click PDB code to view detailed information in PDB database. Alternatively, users can download the PDB file of the polymerase.